Single-Center vs Multi-Center Studies: What We've Learned About Scaling Imaging AI
Reading time /
3 min
Research


From Proof-of-Concept to Scale
Over the past ten years, artificial intelligence (AI) has progressed from research-oriented prototypes to clinical decision support systems in medical imaging. The U.S. Food and Drug Administration (FDA) has approved more than a thousand AI/ML-assisted medical devices, the majority of which are imaging-based.
The early successes of imaging AI often begin in the highly resourceful academic setting. A research team, often affiliated with a leading technical university, collaborates with a renowned academic medical center. The purpose of this collaboration is to leverage a carefully curated retrospective dataset precisely matching the target application of the AI model. These datasets usually have structured reporting, uniform imaging protocols, stable scanner configurations, and relatively complete metadata. In such controlled settings, model development is rapid as datasets are easily standardized and clinical outcomes are well-defined. Internal validation of such AI models demonstrates excellent performance characteristics such as high area under the curve (AUC), sensitivity, and specificity because the training and validation datasets share a common institutional fingerprint.
However, this success may be more a result of intra-institutional optimization than true generalizability. When the same model is deployed in the real world, several destabilizing factors emerge. These include variability in data structure and demographics, imaging acquisition heterogeneity, workflow artifacts and other hidden confounders.1,2
Defining Single-Center vs Multi-Center Imaging Studies
A single-center study derives all data from one institution, often a single academic hospital or health system. Data typically share common characteristics such as uniform scanner fleet (e.g., limited vendors/models), standardized acquisition protocols, consistent reporting language, homogeneous patient referral patterns. Such datasets offer low variance and tight control over confounders. While, a multi-center study aggregates data across two or more independent institutions. These institutions may differ in geography and patient demographics, scanner vendors and software versions. They may also vary in imaging protocols, clinical documentation practices and care delivery settings including academic, community, specialty hospitals.
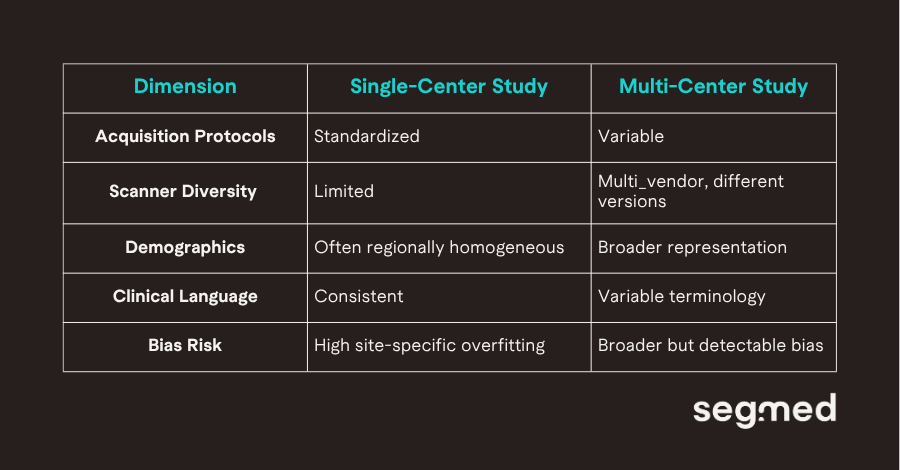
The table above represents the heterogeneity in key structural elements of both of the studies.
The Promise and Pitfalls of Single-Center Studies
Single-center datasets have been the backbone of modern imaging AI development. For early innovation and feasibility studies and proof-of-concept models, such conditions are not only convenient but they are strategically appropriate.
Apart from standardization in data collection and storage, single-center studies offer advantages such as tight collaboration between clinicians and data scientists. From a modeling perspective, single-center AI model development reduces unwanted variance and allows algorithms to identify clinically relevant signals with greater statistical efficiency. Feedback loops between model developers and clinical stakeholders are proactive. Iterative refinements adjusting labeling strategies, recalibrating thresholds, or testing architectural changes can occur without the logistical burden of cross-site coordination. If a tool is intended for deployment within the same institutional ecosystem in which it was trained, performance may likely remain stable.
“Single-center datasets accelerate innovation, but they can also encode institutional fingerprints into AI models.”
The difficulty arises when institutional optimization is mistaken for generalizability. Evidence from cross-hospital evaluations suggests that models trained in homogeneous environments often encode site-specific characteristics rather than purely pathological features. Research published in NPJ Digital Medicine showed that deep learning systems could leverage hospital process artifacts, rather than disease biology to generate predictions. These findings highlight a structural vulnerability: models may internalize contextual signals such as scanner-specific noise patterns, acquisition conventions, or workflow correlates that do not transfer beyond the originating site.
Demographic and technical homogeneity compound the issue. A single institution reflects a specific referral base, socioeconomic distribution, and disease prevalence profile. Scanner vendors, hardware generations, and reconstruction techniques may be limited in such scenarios. Without exposure to broader variability during training, models risk uneven performance across patient subgroups. Disparities in algorithmic performance across race and sex have been documented in imaging AI systems. One such example includes an analysis published in Nature Medicine, underscoring the importance of representative data during development.
Perhaps the most persistent risk is misplaced confidence derived from internal validation. When training and test sets are sampled from the same institution. Many imaging AI studies lack rigorous external validation and may therefore overestimate real-world readiness. Across multiple evaluations, measurable declines in performance have been observed when models are assessed outside their training institution. In practical terms, a model that achieves 0.95 AUC internally may experience meaningful degradation once exposed to new patient populations, scanner ecosystems, and documentation practices. Single-center studies remain essential for innovation, but without external validation, they measure optimization not scalability.
Why Multi-Center Data Changes Everything?
“Multi-center datasets stress-tests AI systems before they reach clinical environments.”
Multi-center data fundamentally reshape how imaging AI systems are trained and evaluated because they introduce structured exposure to real-world heterogeneity. Differences in patient populations, care settings across institutions cause variability in imaging data. Scanner vendors, software versions, field strengths, and reconstruction pipelines are all examples of how images vary due to technical characteristics of different imaging systems. Variability exists even if two centers are using the same scanner model; different acquisition parameters and reconstruction methods can be found due to local protocols or different preferences of radiologists at each center. These seemingly minor discrepancies result in images that have different characteristics than anticipated and can create measurement shifts that aren't apparent during single-center development.
By using multi-center images for training, pathological features that occur consistently across the sites are learned by the cross-site model, rather than by training on site-specific characteristics alone. These advantages lead to improvements in generalizability and stability. Studies involving more than one institution have provided ample evidence that there is usually a decrease in how well models work when they are tested at different institutions from where they were developed.
Declines in model performance when utilized in different settings are often not indicative of a flaw in the model; rather, they are commonly attributable to the distribution shifts a model encounters when being used outside its normal site of development. Through the use of multi-center validation, potential weaknesses are identified early and offer opportunities for recalibrating models prior to implementation in the clinical setting. Broader datasets allow stratified evaluation across demographic groups and care environments, helping to uncover hidden biases that single-center cohorts may obscure. External validation across independent institutions has therefore become central to clinical trust and regulatory confidence.3,4
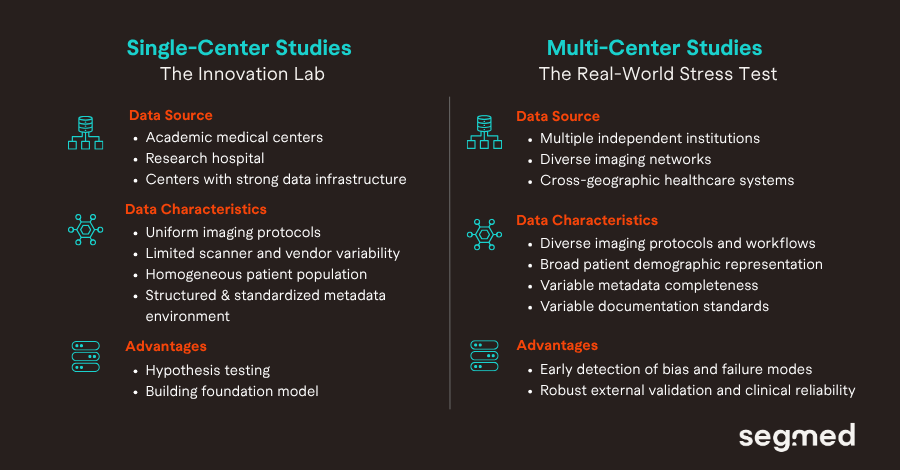
Lessons Learned from Real-World Imaging Data at Scale
Working with globally distributed imaging datasets makes one principle unmistakably clear: heterogeneity is the norm, not the exception. Across institutions and geographies, variability appears in metadata completeness, reporting language, labeling standards, and annotation consistency. Even when imaging pixels appear comparable, the surrounding clinical context often differs. Effective harmonization therefore requires more than technical normalization; it demands clinical insight to distinguish meaningful variation from noise.
This variability creates a natural tension between data complexity and model readiness. Highly heterogeneous datasets introduce statistical noise, may slow early convergence, and often require larger sample sizes or domain adaptation techniques. However, models trained under such conditions tend to be more resilient at deployment. Exposure to diverse scanners, protocols, and patient populations during development reduces reliance on site-specific artifacts and improves robustness under distribution shift.
At scale, it also becomes clear that data strategy must evolve alongside algorithm maturity. Early AI efforts often emphasize dataset volume to establish innovation feasibility. Over time, representativeness becomes more critical than size alone. A structured progression typically strengthens generalizability. In an award-winning study performed by the Segmed team, radiology reporting patterns were examined for oligometastatic disease. Specialty oncology centers demonstrated higher adoption of the term compared with general hospitals.
A single cancer-center study would therefore overestimate prevalence relative to a broader multi-center sample. This underscores a broader reality, institutional culture influences not only imaging acquisition but also diagnostic terminology. AI systems incorporating NLP or structured reporting signals must explicitly account for such heterogeneity to avoid embedding linguistic bias into model outputs.
Conclusion: Data as the Foundation for Scalable Imaging AI
Single-center studies remain indispensable to the advancement of imaging AI. They provide the controlled conditions necessary for hypothesis testing, rapid iteration, foundation model and early performance validation. Yet scalability demands a different benchmark. Multi-center validation is the mechanism through which these become attainable. It reveals whether a model is resilient to distributional changes and whether performance holds across institutions that differ in geography, patient mix, technical infrastructure, and reporting culture. Without such validation, strong internal metrics risk overstating clinical readiness.
As imaging AI matures, the methodological expectation is becoming clearer. Internal validation demonstrates feasibility within a defined environment. External, multi-center validation demonstrates durability across environments. Ultimately, scalable imaging AI is not achieved through algorithmic sophistication alone. Model architecture, training strategies, and computational efficiency matter but they cannot compensate for diverse and unrepresentative data. Robustness emerges from deliberate exposure to real-world heterogeneity during development and validation.
Multi-Center Validation with Segmed: The Path to Scalable AI
With the transition from development to implementation of imaging AI, the capability to validate across various clinical contexts is key. While it may be possible to achieve excellent results using training data sets from a single-center, the real test of scalability is the generalizability across centers, demographics, and imaging modalities.
By leveraging our extensive network of partners, Segmed allows access to large, multi-center imaging data sets, which simulate real-life diversity. By virtue of wide geographical representation and diversified imaging contexts, our data sets can facilitate external validation, bias evaluation, and performance benchmarking of AI models.
Get in touch to learn more about how multi-center imaging data can help you develop AI models that are accurate, robust, and ready for real-world clinical use.
Frequented Asked Questions - F.A.Q.
What is the difference between single-center and multi-center imaging studies?
The difference between single-center and multi-center imaging studies lies in the number of institutions contributing imaging data and how that affects data variability, generalizability, and model robustness.
A single-center study collects imaging data from one hospital or institution.
Key characteristics:
- Uniform scanner vendors and protocols
- Homogeneous patient population
- Consistent acquisition parameters
- Limited demographic diversity
Advantages:
- Controlled environment
- Faster execution
- Lower operational complexity
Limitations:
- High risk of overfitting
- Limited external validity
- Poor generalizability to other healthcare settings
A multi-center study aggregates imaging data from multiple hospitals or research institutions.
Key characteristics:
- Diverse scanner manufacturers
- Variable acquisition protocols
- Broader demographic representation
- Greater real-world variability
Advantages:
- Improved model robustness
- Better generalizability
- Reduced institutional bias
- Higher regulatory credibility
Limitations:
- Increased logistical complexity
- Data harmonization challenges
- Regulatory and data-sharing barriers
Bottom Line
Single-center datasets often fail to capture real-world heterogeneity in clinical imaging. Multi-center imaging studies better reflect real-world deployment conditions and are considered the gold standard for validating AI models in radiology and medical imaging.
Why do AI models perform worse during external validation?
AI models frequently show performance degradation during external validation because they encounter data distributions that differ from the training dataset. This phenomenon is primarily driven by distribution shifts.
Key causes of performance drop include:
- Domain Shift- Differences in scanner vendors, imaging protocols, reconstruction algorithms. Even subtle technical differences can significantly alter pixel intensity distributions.
- Demographic Variation- Training data may underrepresent: ethnic minorities, pediatric or geriatric populations, rare disease subtypes, etc. Models trained on narrow populations fail when applied to broader groups.
- Overfitting to Institutional Artifacts- Models may inadvertently learn site-specific metadata, scanner-specific noise patterns, institutional annotation styles. These features do not generalize externally.
Why is multi-center validation important in medical imaging AI?
Multi-center validation is critical for ensuring robustness, fairness, and regulatory readiness of AI models in medical imaging. Exposure to heterogeneous datasets allows models to learn invariant features, reduce overfitting, and perform reliably across institutions and reflects real-world implementation.
Multi-center validation also boosts clinical and regulatory credibility. Regulatory agencies such as the U.S. Food and Drug Administration and the European Medicines Agency expect evidence of performance across diverse populations before approving AI-based medical devices.
How does dataset bias affect imaging AI performance?
Dataset bias occurs when training data does not accurately represent the target population or real-world variability. This significantly impacts imaging AI performance.
Consequences of Dataset Bias:
- Reduced external validation performance
- Increased false positives/negatives
- Amplification of healthcare disparities
- Regulatory rejection
- Loss of clinician trust
Mitigation Strategies:
- Multi-center data aggregation
- Data harmonization techniques
- Prospective external validation
- Federated learning approaches
References
- Kelly CJ, Karthikesalingam A, Suleyman M, Corrado G, King D. Key Challenges for Delivering Clinical Impact with Artificial Intelligence. BMC Medicine [Internet]. 2019 Oct 29;17(1). Available from: https://bmcmedicine.biomedcentral.com/articles/10.1186/s12916-019-1426-2
- Zech JR, Badgeley MA, Liu M, Costa AB, Titano JJ, Oermann EK. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: A cross-sectional study. Sheikh A, editor. PLOS Medicine [Internet]. 2018 Nov 6;15(11):e1002683. Available from: https://journals.plos.org/plosmedicine/article?id=10.1371/journal.pmed.1002683
- Willemink MJ, Koszek, Wojciech A, Hardell C, Wu J, Fleischmann D, Harvey H, et al. Preparing Medical Imaging Data for Machine Learning. Radiology [Internet]. 2020 [cited 2026 Feb 24];295(1):4–15. Available from: https://doi.org/10.1148/radiol.2020192224
- Roberts M, Driggs D, Thorpe M, Gilbey J, Yeung M, Ursprung S, et al. Common pitfalls and recommendations for using machine learning to detect and prognosticate for COVID19 using chest radiographs and CT scans. Nature Machine Intelligence [Internet]. 2021;3(3):199–217. Available from: https://doi.org/10.1038/s42256021003070


